[Top 5 Things I Wish I Knew When I Started a PhD]
Added on 06/05/2015
In a short moment self-reflection, I made a list of the five most important things that doing research with a lot of data has taught me. And I learned the hard way - wasting a lot of time and energy re-doing things instead of being smart about it at the beginning. Note to self: I should read this once a year or so.
1. Take good notes while reading literature
 There is nothing more annoying than having an excellent set of results to present, but finding it hard to go back to the motivation of the project and writing about why the results were produced. I find it really important to take notes at the beginning of a project, even if I am not sure if they will be relevant when my results are done.
There is nothing more annoying than having an excellent set of results to present, but finding it hard to go back to the motivation of the project and writing about why the results were produced. I find it really important to take notes at the beginning of a project, even if I am not sure if they will be relevant when my results are done.
I save a few notes at the top of each PDF I read and I also save relevant referenced sentences in a separate file. Sometimes it’s even just interesting ideas of other authors that I agree or disagree with.
Since experimentation work usually takes months to produce anything, having those background notes to fall back on is a big help when I want to get a paper ready or when I want to quickly scan through the papers I read. It is also great to read my notes from time to time while I am doing experiments, so that I don’t loose sight of the ‘big picture’.
2. Make scripts. As many as possible.
 Automation of experiments in form of bash scripts is something that we all kind of know about but often don’t bother to do. However, I find that scripts that can set up and run simulations are very helpful in the long run and especially when I come back to re-running experiments I have forgotten details of.
Automation of experiments in form of bash scripts is something that we all kind of know about but often don’t bother to do. However, I find that scripts that can set up and run simulations are very helpful in the long run and especially when I come back to re-running experiments I have forgotten details of.
Making bash scripts takes time though, which is why I didn’t do it at first. It didn’t seem that important. Then I had to change some simulation parameters and re-run everything. Again and again. Now, I simply invest the necessary time at the beginning, which makes me much more comfortable and ready to re-run anything I need to. And when I say script everything I mean everything - from automating the way input files are set up, executable files are run, results are sorted, sometimes even the way I move between directories during experimentation.
3. Take good notes of the experiments performed
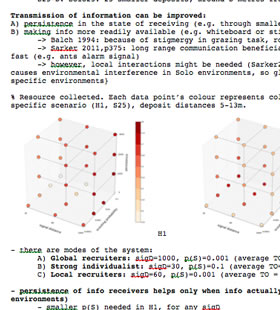 I use simulations instead of running real-world experiments because I can do tens of runs in a matter of minutes. Simulations are especially useful for experimenting with parameters. Vary this parameter, vary that. It’s all perfectly clear while I am doing it.
I use simulations instead of running real-world experiments because I can do tens of runs in a matter of minutes. Simulations are especially useful for experimenting with parameters. Vary this parameter, vary that. It’s all perfectly clear while I am doing it.
I used to sort my results with different parameters into folders and name the folders with abbreviations of parameter names and values. Until I had a big experiment that lasted for about half a year and then wanted to come back to it in a year. I understood nothing.
Now, I try to take very detailed notes of what experiments I am running and why. I note down what worked and what didn’t and why I decided to go with a particular set of parameters. Even if I delete data because it doesn’t produce anything useful, I note that down so that I don’t wander down that particular avenue in the future. Most importantly, I avoid abbreviations. Yes, my folder names are usually over 100 characters long, but at least I understand exactly what they mean.
4. Refactor, inherit, comment
 Good code means well commented code with a minimum amount of lines to do the job asked. It’s generally known that making your code readable is a must if you ever want to come back to your experiments. In Object Oriented Programming, this also includes subclassing rather than using If statements when a particular algorithm gets too complicated.
Good code means well commented code with a minimum amount of lines to do the job asked. It’s generally known that making your code readable is a must if you ever want to come back to your experiments. In Object Oriented Programming, this also includes subclassing rather than using If statements when a particular algorithm gets too complicated.
Comments are at least as important as the structure of the code itself. For example, I find it a good practise to describe important variables when they are declared. Or to explain why a function (and especially a mathematical function that computes a result from variables) does that it does.
5. Make production of final figures as painless as possible
 After a paper is submitted, it usually comes back from peer review with a number of requested changes. But the peer review process can take months. After so long, I often find it hard to figure out how exactly I produced particular figures from my data when I need to change them, especially as I usually have a lot of data and a lot of figures produced during analysis.
After a paper is submitted, it usually comes back from peer review with a number of requested changes. But the peer review process can take months. After so long, I often find it hard to figure out how exactly I produced particular figures from my data when I need to change them, especially as I usually have a lot of data and a lot of figures produced during analysis.
To minimise the time I need to reproduce the figures that appear in the paper, I now do two things: 1. Produce figures with a script (I use python) and setup some way of quickly producing just the figures for the paper. E.g. setting up a global variable in the python script, that if set to true will call the relevant functions, is a good idea. 2. Either output those important figures in a separate folder, or make aliases (that’s ‘shortcuts’, Windows users) to them in a separate folder. This saves a lot of time I previously used to spend going through image files when I looked for the updated images.
It’s not all just about what you do..
Everyone is different in the way they function. Just like many people, I was conditioned by a 9-5 job for a while (well more like 9-8 but ok). Doing a PhD has been a great opportunity to find the actual ideal time for me to work. Ideal working hours and work / rest balance are important. I guess, until you figure out the way your own brain works, you will work against yourself to some extent. Of course, if you work with others on a day-to-day basis, this becomes more difficult.











{Please enable JavaScript in order to post comments}